Cunoașterea predicției unui model este utilă. Știind cât de încrezător este acea predicție, cu atât mai mult. Predicția conformă oferă exact asta: intervale de predicție valide statistic cu acoperire garantată (în anumite condiții), indiferent de modelul de bază sau de distribuția datelor.
În această postare, împerechem două instrumente puternice: TabPFNa transformator preantrenat pentru date tabulareși nnetsaucelui PredictionInterval (care implementează Split Conformal Prediction), care include orice regresor compatibil cu scikit-learn într-un predictor conform. Demonstrăm întreaga conductă pe setul de date despre diabet, mai întâi în Python, apoi în R prin reticulat. Ambele versiuni produc rezultate identice: o rată de acoperire de 96,7% la un nivel nominal de 95%.
!pip install tabpfn tabpfn_client
!pip install nnetsauce
import tabpfn_client
API_TOKEN = "" # <- Paste your TabPFN token here (from https://priorlabs.ai/tabpfn)
tabpfn_client.set_access_token(API_TOKEN)
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split
from tabpfn_client import TabPFNRegressor
from sklearn.metrics import mean_squared_error
import numpy as np
reg = TabPFNRegressor()
X, y = load_diabetes(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
reg.fit(X_train, y_train)
preds = reg.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds))
print(-rmse)
00:00 Fitting... |
WARNING:tabpfn_client.client:The provided train set hashes match previously uploaded train sets.
00:00 Fitting... Done!
00:00 Predicting... -
WARNING:tabpfn_client.client:The provided test set hash matches a previously uploaded test set.
00:01 Predicting... Done!
-51.559912022529886
import nnetsauce as ns
reg_conformal = ns.PredictionInterval(reg, level=95)
reg_conformal.fit(X_train, y_train)
preds = reg_conformal.predict(X_test, return_pi=True)
00:00 Fitting... |
WARNING:tabpfn_client.client:The provided train set hashes match previously uploaded train sets.
00:00 Fitting... Done!
00:00 Predicting... -
WARNING:tabpfn_client.client:The provided test set hash matches a previously uploaded test set.
00:01 Predicting... Done!
00:00 Predicting... -
WARNING:tabpfn_client.client:The provided test set hash matches a previously uploaded test set.
00:01 Predicting... Done!
00:00 Predicting... -
WARNING:tabpfn_client.client:The provided test set hash matches a previously uploaded test set.
00:01 Predicting... Done!
print(f"coverage_rate: {np.mean((preds.lower<=y_test)*(preds.upper>=y_test))}")
coverage_rate: 0.9662921348314607
import warnings
import matplotlib.pyplot as plt
warnings.filterwarnings('ignore')
split_color="green"
split_color2 = 'orange'
local_color="gray"
def plot_func(x,
y,
y_u=None,
y_l=None,
pred=None,
shade_color="",
method_name="",
title=""):
fig = plt.figure()
plt.plot(x, y, 'k.', alpha=.3, markersize=10,
fillstyle="full", label=u'Test set observations')
if (y_u is not None) and (y_l is not None):
plt.fill(np.concatenate((x, x(::-1))),
np.concatenate((y_u, y_l(::-1))),
alpha=.3, fc=shade_color, ec="None",
label = method_name + ' Prediction interval')
if pred is not None:
plt.plot(x, pred, 'k--', lw=2, alpha=0.9,
label=u'Predicted value')
#plt.ylim((-2.5, 7))
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.legend(loc="upper right")
plt.title(title)
plt.show()
max_idx = 50
plot_func(x = range(max_idx),
y = y_test(0:max_idx),
y_u = preds.upper(0:max_idx),
y_l = preds.lower(0:max_idx),
pred = preds.mean(0:max_idx),
shade_color=split_color2,
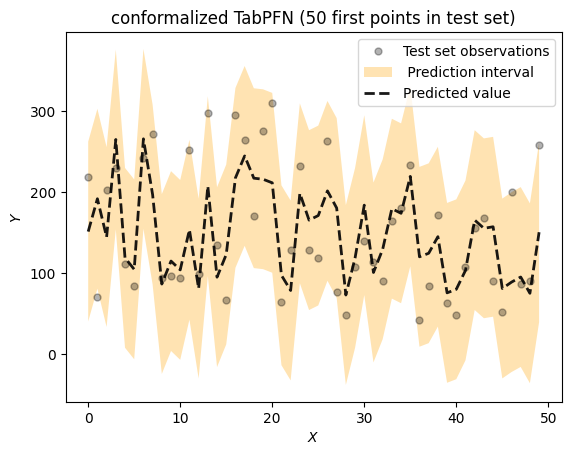
title = f"conformalized TabPFN ({max_idx} first points in test set)")
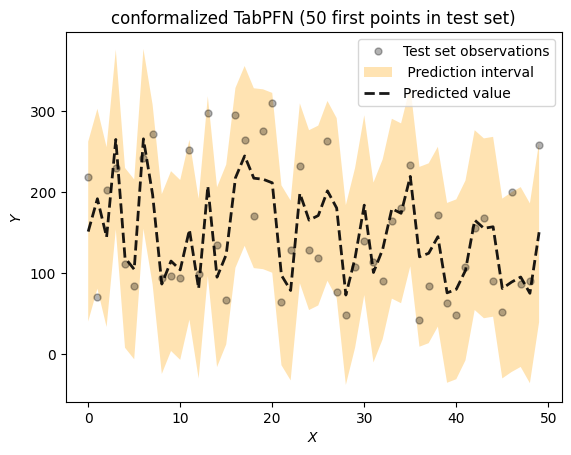
Pentru această versiune R, am folosit R în același notebook ca Python, în Google Colab.
%load_ext rpy2.ipython
%R install.packages("reticulate")
%%R
# Conformalized TabPFN in R via reticulate
library(reticulate)
# ── 0. Python environment ──────────────────────────────────────────────────────
# Use your preferred Python env. Uncomment one (automatic on Google Colab):
# use_python("/usr/bin/python3")
# use_virtualenv("r-tabpfn")
# use_condaenv("r-tabpfn")
# Install required packages into the active Python env (run once)
# py_install(c("tabpfn", "tabpfn_client", "nnetsauce", "scikit-learn",
# "matplotlib", "numpy"), pip = TRUE)
# ── 1. Imports ─────────────────────────────────────────────────────────────────
sklearn_datasets <- import("sklearn.datasets")
sklearn_model_sel <- import("sklearn.model_selection")
sklearn_metrics <- import("sklearn.metrics")
tabpfn_client <- import("tabpfn_client")
ns <- import("nnetsauce")
np <- import("numpy")
plt <- import("matplotlib.pyplot")
warnings <- import("warnings")
# ── 2. TabPFN API token ────────────────────────────────────────────────────────
API_TOKEN <- "" # <-- paste your TabPFN token here (from https://priorlabs.ai/tabpfn)
tabpfn_client$set_access_token(API_TOKEN)
TabPFNRegressor <- tabpfn_client$TabPFNRegressor
# ── 3. Data ────────────────────────────────────────────────────────────────────
diabetes <- sklearn_datasets$load_diabetes(return_X_y = TRUE)
X <- diabetes((1))
y <- diabetes((2))
split <- sklearn_model_sel$train_test_split(X, y, test_size = 0.2, random_state = 42L)
X_train <- split((1))
X_test <- split((2))
y_train <- split((3))
y_test <- split((4))
# ── 4. Fit TabPFN regressor ────────────────────────────────────────────────────
reg <- TabPFNRegressor()
reg$fit(X_train, y_train)
preds_plain <- reg$predict(X_test)
rmse <- sqrt(sklearn_metrics$mean_squared_error(y_test, preds_plain))
cat(sprintf("TabPFN RMSE: %.4fn", rmse))
# ── 5. Conformal prediction with nnetsauce ─────────────────────────────────────
reg_conformal <- ns$PredictionInterval(reg, level = 95L)
reg_conformal$fit(X_train, y_train)
preds <- reg_conformal$predict(X_test, return_pi = TRUE)
coverage <- np$mean((preds$lower <= y_test) * (preds$upper >= y_test))
cat(sprintf("Coverage rate: %.4fn", coverage))
# ── 6. Plot (first 50 test points) ────────────────────────────────────────────
warnings$filterwarnings("ignore")
max_idx <- 50L
x_range <- np$array(0:(max_idx - 1)) # numeric index
y_obs <- y_test(1:max_idx)
y_upper <- preds$upper(1:max_idx)
y_lower <- preds$lower(1:max_idx)
y_pred <- preds$mean(1:max_idx)
# Build the filled polygon (matplotlib-style concatenation)
x_fill <- np$concatenate(list(x_range, x_range(max_idx:1)))
y_fill <- np$concatenate(list(y_upper, y_lower(max_idx:1)))
fig <- plt$figure()
plt$plot(x_range, y_obs, "k.", alpha = 0.3, markersize = 10L,
label = "Test set observations")
plt$fill(x_fill, y_fill, alpha = 0.3, fc = "orange", ec = "None",
label = "Conformal Prediction interval")
plt$plot(x_range, y_pred, "k--", lw = 2L, alpha = 0.9,
label = "Predicted value")
plt$xlabel("Index")
plt$ylabel("Y")
plt$legend(loc = "upper right")
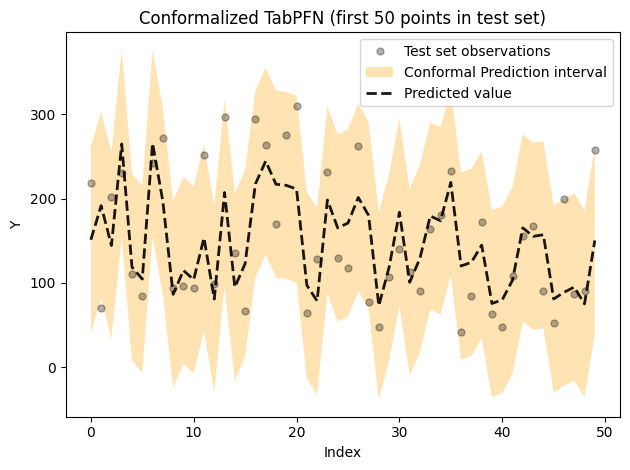
plt$title(sprintf("Conformalized TabPFN (first %d points in test set)", max_idx))
plt$tight_layout()
plt$show()
# To save instead: plt$savefig("conformalized_tabpfn.png", dpi = 150L)
00:02 Fitting... Done!
00:02 Predicting... Done!
TabPFN RMSE: 51.5599
00:01 Fitting... Done!
00:02 Predicting... Done!
00:00 Predicting... -
WARNING:tabpfn_client.client:The provided test set hash matches a previously uploaded test set.
00:01 Predicting... Done!
00:02 Predicting... Done!
Coverage rate: 0.9663